Bloodstream infections (BSIs) are serious infections associated with high rates of morbidity and mortality. Every hour delay in initiation of an effective antibiotic increases mortality due to sepsis by 7%. Turnaround time (TAT) for conventional blood cultures takes 48h, forcing physicians to streamline therapy by exposing patients to broad-spectrum antimicrobials. Our objective was (1) to evaluate the accuracy and TAT of an optimized workflow combining direct matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS) and in-house real-time polymerase chain reaction (PCR) for bacterial identification and antimicrobial resistance profiling directly from positive blood bottles for diagnosing bloodstream infections and (2) to verify the effect of reporting results to medical staff. A total of 103 BSI episodes from 91 patients admitted to three hospitals in São Paulo, Brazil were included. TAT from molecular versus conventional methods was measured and compared. Our protocol showed an overall agreement of 93.5% for genus and 78.5% for species identification; 74.2% for methicillin resistance detection, 89.2% for extended-spectrum β-lactamase profiling, 77.8% for metallo-β-lactamase profiling, and 100% for carbapenemase profile and vancomycin-resistance detection when compared with conventional testing. TAT of molecular sample processing according to our protocol was 38h shorter than conventional methods. Antimicrobial interventions were possible in 27 BSI episodes. Antimicrobial discontinuation was achieved in 12 BSI episodes while escalation of therapy occurred in 15 episodes. Antimicrobial therapy was inadequate in three (12%) BSI episodes diagnosed using results of molecular testing. Our in-house rapid protocol for identifying both bacteria and antimicrobial resistance provided rapid and accurate results, having good agreement with conventional testing results. These results could contribute to faster antimicrobial therapy interventions in BSI episodes.
Bloodstream infections (BSIs) are serious infections associated with high rates of morbidity and mortality among hospitalized patients despite advances in antimicrobial therapy and supportive care.1,2 The consequences on patient care is immense; BSI increases death rate, extends hospital stays in specialized facilities such as intensive care units (ICU), and result in significant additional expenses.1–7
Rapid diagnosis and treatment of BSIs is critical. Mortality due to sepsis, which is a complication of BSI, has been suggested to increase by 7% for every hour of delay in the administration of appropriate antibiotic therapy.8 The gold standard for detecting circulating microorganisms is blood culture (BC) in fluid media. The time required to perform a BC and subsequently identify the causative pathogen usually takes around 48h.9 Delays in microbiological identification force physicians to streamline therapy resulting in excessive patient exposure to broad-spectrum antimicrobials with subsequent risk of developing antibiotic resistance.10
Molecular approaches such as real-time PCR, fluorescence in situ hybridization, DNA sequencing, and more recently, matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS), have been used to expedite pathogen identification.11–15 However, these techniques have limitations, specifically, the high cost and the need for laboratory staff with technical expertise in each method.15,16
These improvements in clinical microbiology testing led to shortening of turnaround time (TAT) for blood culture processing which represents an important factor with potential for improving patient treatment and recovery.17,18 Additionally, antimicrobial stewardship programs are able to use the results of rapid diagnostic tests to increase the efficacy of antibiotic therapy, while minimizing toxicity and decreasing the overall cost of care.19,20 Studies evaluating the clinical impact of rapid blood culture results have been limited by observational study designs and use of historical controls.21,22 Prior studies have evaluated effective interventions after their implementation by antimicrobial stewardship teams.23–25
The objectives of this study were: (1) to determine the accuracy and TAT of a combined MALDI-TOF/real-time PCR workflow for species identification and detection of resistance determinants directly from positive blood cultures bottles in comparison to automated biochemical profiling and phenotypic susceptibility testing from conventional blood culture fluid subcultures, and (2) to determine the effect of reporting results from rapid testing on antimicrobial treatment, especially in relation to intra-laboratory TAT needed to obtain the final result.
Materials and methodsStudy designThis was a prospective study conducted from September 2015 to September 2016 involving 91 patients admitted to three specialty hospitals associated with the Federal University of São Paulo, Brazil: 1, Hypertension and Kidney Hospital (Hospital do Rim e Hipertensão); 2, Pediatric Oncology Institute (Instituto de Oncologia Pediátrica); and 3, Hospital São Paulo. These hospitals have independent administration, bed management, and infection control; they only share a microbiology laboratory located at Hospital São Paulo.
These centers were selected because of the complexity of their patient population. Hypertension and Kidney Hospital admits patients with chronic kidney and cardiovascular diseases, who may or may not have had a solid organ transplant. Pediatric Oncology Institute serves pediatric cancer patients and performs organ and bone marrow transplants. Hospital São Paulo is a general teaching hospital but only patients admitted to the adult oncology and hematology unit were included in this study.
This study was approved by the Research Ethics Committee of the Federal University of São Paulo (Universidade Federal de São Paulo) – UNIFESP, study number 964.693.
Convenience sampling was selected to accommodate a normal molecular laboratory routine. Each work day (Monday through Friday) at 11:59, from the pool of blood culture bottles reported positive during the last 24h with Gram staining results available collected from patients with new BSI episodes admitted to the participating wards, one vial per episode was randomly selected, excluding yeast cultures, and subjected to direct MALDI-TOF identification. Successfully identified samples underwent resistance gene profiling by real-time PCR. Results from samples with conclusive PCR resistance gene profiles were communicated to the medical staff who provided patient information and feedback on subsequent treatment decisions.
Microbiology workflowBlood was cultured using the BACTEC system (BACTEC plus Aerobic/F and Peds Plus/F blood culture media; Becton Dickinson Microbiology Systems, Cockeysville, MD, USA) and continuously monitored for growth in presence of CO2. Gram staining followed by microscopic analyses were performed for positive samples detected by the instrument and partial results were reported to the hospital. Primary cultures testing positive for bacterial growth were routinely subcultured onto blood agar, MacConkey agar, and chocolate agar plates which were then incubated overnight at 37°C and results were immediately reported. At the end of 5-day incubation period, blood cultures were reported as negative if no microorganism had grown by this method.
Bacterial identification and antimicrobial susceptibility tests were performed using a Phoenix instrument (Becton Dickinson Microbiology Systems, Cockeysville, MD, USA) utilizing the panels PMID-121 for Gram negative isolates, PMID-123 for non-fermenters Gram negative bacilli and PMID-104 for Gram positive cocci. Antimicrobial susceptibility test results were provided according to the Clinical & Laboratory Standards Institute M100 document (2016). Prior and during this study, the microbiology laboratory did not have a MALDI-TOF instrument available for routine tests.
After phenotypic processing, bacterial isolates were sent to the Special Laboratory of Clinical Microbiology (Laboratório Especial de Microbiologia Clínica [LEMC]) for long term storage at −20°C in their frozen biobank which is available to university researchers.
Optimized protocol workflowAt the collection time (11:59am) samples were selected and sent to two different laboratories for molecular analyses: microbial identification was performed at the Association Fund for Research Incentives laboratory (Associação Fundo de Incentivo à Pesquisa laboratory) and real-time PCR was performed at LEMC. Both laboratories are located a few meters away from each other and from participating hospitals.
For bacterial identification, a 5mL aliquot was prepared following a previously described protocol.26 Samples were identified by direct MALDI-TOF MS using the VITEK-MS system (bioMérieux, Marcy-l’Etoile, France) with the bioMérieux platform MylaTM v2.0. The software calculated confidence values for each of the two replicates of the tested strains. According to the manufacturer, values between 60.0 and 99.9 indicated a reliable discrimination among species or species group.
For the in-house real-time PCR analysis, 0.5mL of blood was added to 500μL of a commercial solution of phenol and guanidine isothiocynate (Brazol, LGC, Brazil) in a 2-mL screw cap microtube for DNA extraction and vortexed for 3–5s. One hundred and eighty μL of chloroform (8°C) was then added and the sample vortexed for 3–5s. The microtube was centrifuged at 13,000rpm for 12min at 8°C. The supernatant was withdrawn and transferred to a 2-mL microtube containing 500μL of absolute ethanol at 8°C. After vortexing for 3–5s, the mixture was then centrifuged at 13,000rpm for 15min at 8°C. Again the supernatant was removed and washed with 500μL of ethanol (8°C) and then centrifuged at 13,000rpm for 12min at 8°C. Finally, the supernatant was removed and the pellet was allowed to dry at room temperature, then solubilized in 50μL of UltraPure water (Invitrogen, Carlsbad, CA, USA) and incubated at 65°C for 30min.
After DNA extraction, samples were subjected to real-time PCR to look for genes encoding antimicrobial resistance. Toward that end, the result of the bacterial identification by MALDI-TOF was used as guide and the reactions were performed on a Rotor-Gene Q instrument using Rotor-Gene Q Software version 2.1.0 (Qiagen, Hilden, Germany).
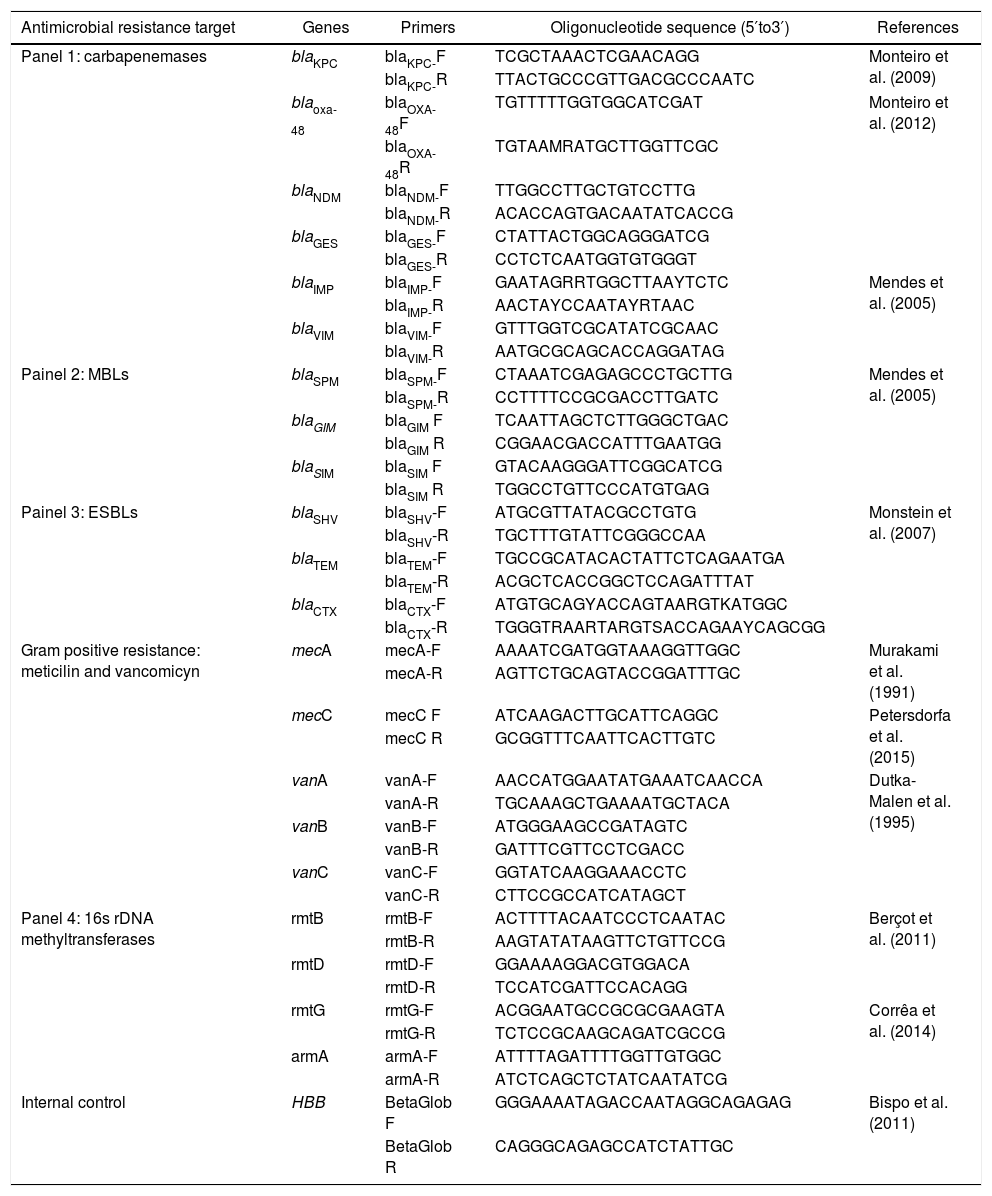
Samples with Gram positive microorganisms were subjected to single real-time PCR reactions for detection of the following genes: mecA, mecC, vanA, vanB, and vanC. For the Gram negative samples, five different gene panels were tested, including carbapenemase, Amber class B metallo-β-lactamase (MBL) and extended-spectrum β-lactamase (ESBL), and 16S rDNA methyltransferase encoding genes. The genes included in these panels were selected based on the local epidemiology of BSI in Brazil. A single assay for the detection of the human β-globin gene (HBB) was used as a blood DNA extraction control. A list of the panels and primers used in this study is shown in Table 1.
Oligonucleotides used in the developed protocol for antimicrobial resistance genes and internal DNA control detection.
| Antimicrobial resistance target | Genes | Primers | Oligonucleotide sequence (5′to3′) | References |
|---|---|---|---|---|
| Panel 1: carbapenemases | blaKPC | blaKPC-F | TCGCTAAACTCGAACAGG | Monteiro et al. (2009) |
| blaKPC-R | TTACTGCCCGTTGACGCCCAATC | |||
| blaoxa-48 | blaOXA-48F | TGTTTTTGGTGGCATCGAT | Monteiro et al. (2012) | |
| blaOXA-48R | TGTAAMRATGCTTGGTTCGC | |||
| blaNDM | blaNDM-F | TTGGCCTTGCTGTCCTTG | ||
| blaNDM-R | ACACCAGTGACAATATCACCG | |||
| blaGES | blaGES-F | CTATTACTGGCAGGGATCG | ||
| blaGES-R | CCTCTCAATGGTGTGGGT | |||
| blaIMP | blaIMP-F | GAATAGRRTGGCTTAAYTCTC | Mendes et al. (2005) | |
| blaIMP-R | AACTAYCCAATAYRTAAC | |||
| blaVIM | blaVIM-F | GTTTGGTCGCATATCGCAAC | ||
| blaVIM-R | AATGCGCAGCACCAGGATAG | |||
| Painel 2: MBLs | blaSPM | blaSPM-F | CTAAATCGAGAGCCCTGCTTG | Mendes et al. (2005) |
| blaSPM-R | CCTTTTCCGCGACCTTGATC | |||
| blaGIM | blaGIM F | TCAATTAGCTCTTGGGCTGAC | ||
| blaGIM R | CGGAACGACCATTTGAATGG | |||
| blaSIM | blaSIM F | GTACAAGGGATTCGGCATCG | ||
| blaSIM R | TGGCCTGTTCCCATGTGAG | |||
| Painel 3: ESBLs | blaSHV | blaSHV-F | ATGCGTTATACGCCTGTG | Monstein et al. (2007) |
| blaSHV-R | TGCTTTGTATTCGGGCCAA | |||
| blaTEM | blaTEM-F | TGCCGCATACACTATTCTCAGAATGA | ||
| blaTEM-R | ACGCTCACCGGCTCCAGATTTAT | |||
| blaCTX | blaCTX-F | ATGTGCAGYACCAGTAARGTKATGGC | ||
| blaCTX-R | TGGGTRAARTARGTSACCAGAAYCAGCGG | |||
| Gram positive resistance: meticilin and vancomicyn | mecA | mecA-F | AAAATCGATGGTAAAGGTTGGC | Murakami et al. (1991) |
| mecA-R | AGTTCTGCAGTACCGGATTTGC | |||
| mecC | mecC F | ATCAAGACTTGCATTCAGGC | Petersdorfa et al. (2015) | |
| mecC R | GCGGTTTCAATTCACTTGTC | |||
| vanA | vanA-F | AACCATGGAATATGAAATCAACCA | Dutka-Malen et al. (1995) | |
| vanA-R | TGCAAAGCTGAAAATGCTACA | |||
| vanB | vanB-F | ATGGGAAGCCGATAGTC | ||
| vanB-R | GATTTCGTTCCTCGACC | |||
| vanC | vanC-F | GGTATCAAGGAAACCTC | ||
| vanC-R | CTTCCGCCATCATAGCT | |||
| Panel 4: 16s rDNA methyltransferases | rmtB | rmtB-F | ACTTTTACAATCCCTCAATAC | Berçot et al. (2011) |
| rmtB-R | AAGTATATAAGTTCTGTTCCG | |||
| rmtD | rmtD-F | GGAAAAGGACGTGGACA | ||
| rmtD-R | TCCATCGATTCCACAGG | |||
| rmtG | rmtG-F | ACGGAATGCCGCGCGAAGTA | Corrêa et al. (2014) | |
| rmtG-R | TCTCCGCAAGCAGATCGCCG | |||
| armA | armA-F | ATTTTAGATTTTGGTTGTGGC | ||
| armA-R | ATCTCAGCTCTATCAATATCG | |||
| Internal control | HBB | BetaGlob F | GGGAAAATAGACCAATAGGCAGAGAG | Bispo et al. (2011) |
| BetaGlob R | CAGGGCAGAGCCATCTATTGC |
The TAT for processing positive blood cultures was divided into three components. Time to positivity (TTP): corresponds to the incubation time necessary for microbial growth in the BACTEC system to determine positivity; TAT1: time between determination of culture positivity and reporting of isolate identification and antibiotic resistance results using molecular testing; and TAT2: time between determination of culture positivity and reporting of isolate identification and antibiotic resistance results using conventional microbiological methods.
TAT1 and TAT2 times were calculated and the median for each was considered the expected amount of time to generate a report using either conventional BC processing and biochemical identification or the developed optimized protocol.
Clinical data acquisitionThe medical group members consisted of four infectious disease physicians from the three participating centers and their teams. The medical groups from each center received real-time messaging of validated results with the rapid (optimized) protocol through a mobile app. Antibiotic recommendations were not provided by the testing laboratory; antimicrobial therapy was decided by each medical group based on the guidelines of each participating hospital.
The following baseline data were collected from each patient upon enrollment: patient demographics (e.g., age and sex); major comorbid conditions; immune status (neutropenia); site of suspected or confirmed infection; surgery/procedures for suspected site of infection prior to enrollment, such as renal or bone marrow transplantation; and survival status at discharge.
A BSI episode was defined as the isolation of a bacterial or fungal pathogen from at least one BC. A BSI episode was considered new if it was detected at least 14 days from the previous infection with positive blood culture or if a different microorganism was isolated in a new blood sample. Neutropenia was defined as an absolute neutrophil count of <500cells/mm3 within 48h of onset of bacteremia for children or <1500cells/mm3 for adults.
Patients without any identifiable source of infection were classified as having primary bacteremia. Secondary BSIs were defined as those with a clear source of bacteremia other than a central line. Sources of secondary BSIs were identified using culture of samples obtained from distant sites that yielded the same pathogen with an identical antibiotic resistance pattern. Distant sites were defined as sites other than a central line where an infection was diagnosed (e.g., pneumonia, urinary tract infection, or abdominal infection).
Empirical antimicrobial therapy was initiated according to the standard protocol of each center at the time a patient presented with fever and suspicion of BSI, and immediately after blood collection for culture. Antimicrobial therapy was analyzed and assessed by the judgment of medical staff in two periods: Period 1: antibiotics in use prior study enrollment; Period 2: antibiotics in use after the rapid protocol result was reported to the physicians. The treatment was considered inadequate when the antibiotic regimen was not indicated for the microorganism identified or with resistance mechanism detected in the molecular test.
The medical group leader was asked to indicate whether the results from the molecular protocol were used to support antimicrobial therapy (yes or no) and if treatment modifications (drug escalation or de-escalation) were prescribed to the patient based on the molecular protocol results retrospectively by filling a patient's clinical database.
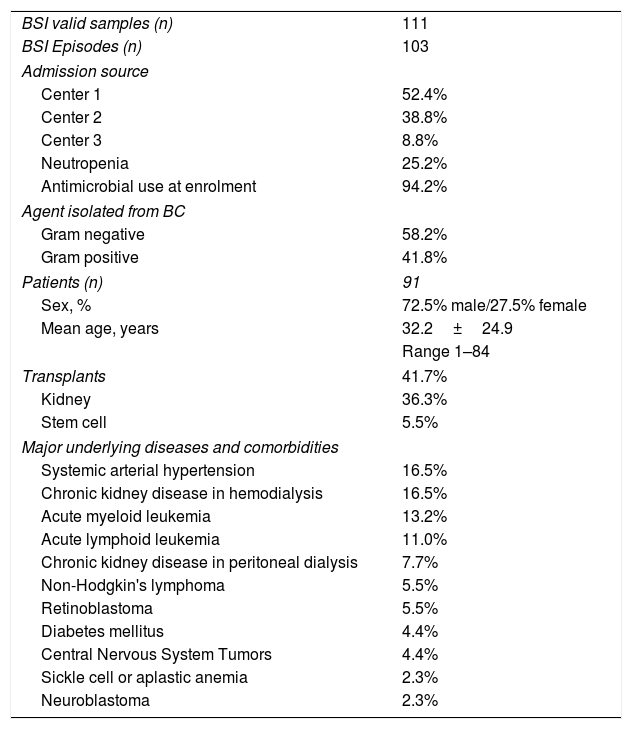
ResultsStudy samples and patient characteristicsDuring the study period, a total of 166 BSI samples were selected. Fifty-five samples were excluded: 34 yielded inconclusive results using MALDI-TOF MS, seven tested negative or inconclusive using real-time PCR, three were collected from organ donors (not hospitalized), and 11 samples did not have clinical data available. Results were validated for 111 samples corresponding to 103 BSI episodes from 91 patients and immediately reported to the appropriate medical group. Samples were collected from peripheral access (n=68) or central line catheter (n=43). Patient demographics and samples included in this study are described in Table 2.
Patients demographics, samples and bloodstream infection (BSI) episodes included in the study.
| BSI valid samples (n) | 111 |
| BSI Episodes (n) | 103 |
| Admission source | |
| Center 1 | 52.4% |
| Center 2 | 38.8% |
| Center 3 | 8.8% |
| Neutropenia | 25.2% |
| Antimicrobial use at enrolment | 94.2% |
| Agent isolated from BC | |
| Gram negative | 58.2% |
| Gram positive | 41.8% |
| Patients (n) | 91 |
| Sex, % | 72.5% male/27.5% female |
| Mean age, years | 32.2±24.9 |
| Range 1–84 | |
| Transplants | 41.7% |
| Kidney | 36.3% |
| Stem cell | 5.5% |
| Major underlying diseases and comorbidities | |
| Systemic arterial hypertension | 16.5% |
| Chronic kidney disease in hemodialysis | 16.5% |
| Acute myeloid leukemia | 13.2% |
| Acute lymphoid leukemia | 11.0% |
| Chronic kidney disease in peritoneal dialysis | 7.7% |
| Non-Hodgkin's lymphoma | 5.5% |
| Retinoblastoma | 5.5% |
| Diabetes mellitus | 4.4% |
| Central Nervous System Tumors | 4.4% |
| Sickle cell or aplastic anemia | 2.3% |
| Neuroblastoma | 2.3% |
Primary bacteremia was identified in 75 (72.8%) BSI episodes, 52 of which were associated with a central line catheter. Sources of secondary BSIs (27.2%) were: urinary tract (13), gastrointestinal tract (8), skin (3), lungs (2), and others (2). In 26 (25.2%) BSI episodes, the patients were neutropenic; 38 BSI patients had been transplanted. Seven (7.7%) patients died during the study period, and five (5.5%) BSI episodes were considered new BSIs as they occurred within 14 d from the original BSI. Underlying diseases and comorbidities are described in Table 2.
Optimized vs. conventional microbiological workflowMALDI-TOF MS was performed taking samples directly from BC bottles of 103 BSI episodes; 33 different microorganisms were identified. Compared with conventional BC processing, the optimized method achieved correct genus identification in 100 episodes and correct genus and species identification in 84 episodes, having 93.5% and 78.5% agreement with conventional microbiological methods, respectively.
A 74.2% categorical agreement was found by the in-house real-time PCR testing for methicillin resistance in Gram positive cocci as compared with the standard method. Among 31 tested staphylococcal samples, 14 mecA positive samples grew isolates that were also found methicillin resistant by phenotypic testing, while six methicillin-resistant samples tested negative for mecA/C and two susceptible samples tested positive mecA. Nine methicillin susceptible isolates were recovered from mecA/C negative samples). A 100% agreement was observed for vancomycin-resistant Enterococci strains, with the vanA gene reported in one Enterococcus faecium sample.
Among Enterobacteriaceae, we found an 89.2% categorical agreement between the phenotypic method for 3rd generation cephalosporin resistance profiling and ESBL molecular detection and 100% for carbapenem-resistance profiling. Nine samples tested positive for blaKPC. MBL-encoding genes were not detected by real-time PCR while the standard method detected four carbapenem-resistant non-fermentative rods, with a 77.8% agreement between the methods. Aminoglycoside resistance was identified in three samples: one Pseudomonas aeruginosa (rmtD) and two Klebsiella pneumoniae (rmtG) isolates. The K. pneumonaiae samples showed a pan-resistance profile when the rmtG gene was co-associated with blaKPC and blaCTX, or blaTEM.
Turnaround timeThe mean TTP of the 111 samples was 14.2h. Using conventional microbiological techniques, the average time to complete and report Gram stain identification and antimicrobial susceptibility testing was 48.2h.
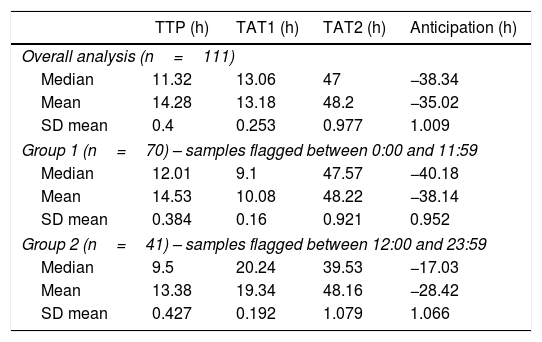
Using the optimized protocol following Gram stain identification, time to organism identification and resistance determination was 13.2h on average, which was approximately 35h faster than conventional methods. TAT components and expected time for obtaining final results are reported in Table 3.
Intra-laboratory turnaround time (TAT) components for molecular and phenotypic analysis of 111 samples included in this study.
| TTP (h) | TAT1 (h) | TAT2 (h) | Anticipation (h) | |
|---|---|---|---|---|
| Overall analysis (n=111) | ||||
| Median | 11.32 | 13.06 | 47 | −38.34 |
| Mean | 14.28 | 13.18 | 48.2 | −35.02 |
| SD mean | 0.4 | 0.253 | 0.977 | 1.009 |
| Group 1 (n=70) – samples flagged between 0:00 and 11:59 | ||||
| Median | 12.01 | 9.1 | 47.57 | −40.18 |
| Mean | 14.53 | 10.08 | 48.22 | −38.14 |
| SD mean | 0.384 | 0.16 | 0.921 | 0.952 |
| Group 2 (n=41) – samples flagged between 12:00 and 23:59 | ||||
| Median | 9.5 | 20.24 | 39.53 | −17.03 |
| Mean | 13.38 | 19.34 | 48.16 | −28.42 |
| SD mean | 0.427 | 0.192 | 1.079 | 1.066 |
TTP, time to positivity; TAT1, timing for molecular report; TAT2, timing for conventional microbiology final result.
The samples were separated into two groups to better understand the performance of the developed protocol. Among the 111 BSI samples, 70 were flagged as culture positive and optimized protocol was performed on the same day (between 00:00 and 11:59, 24h clock), these samples were placed in group 1. Forty-one samples were flagged as culture positive later in the day (between 12:00 and 23:59, 24h clock) and were thus included in the molecular testing for the following day, these samples were placed in group 2. As expected, group 1 report was more promptly made available to physicians and with a time advantage as compared to phenotypic testing (40.2 group 1 vs 17.0h group 2). Phenotypic BC processing time did not show discrepancy in the results between the two groups.
Antimicrobial utility and adequacyIn 97 (94.2%) BSI episodes, patients were on antibiotic treatment at the time blood cultures tested positive. Forty-one (39.8%) patients were on vancomycin and 15 (15.6%) patients were on polymyxin B.
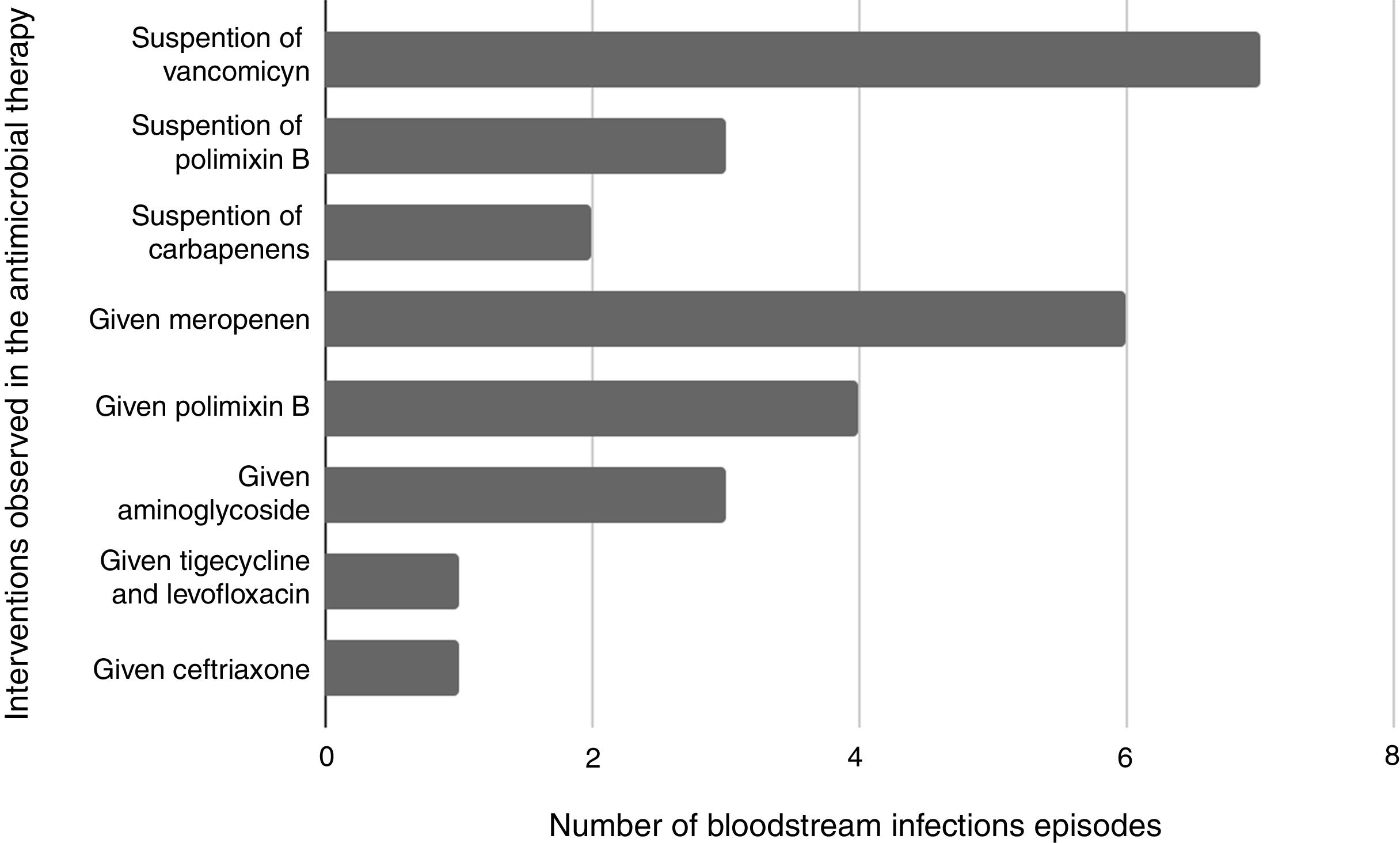
Antibiotic interventions were possible in 27 (26.2%) BSI episodes included in this study. Most interventions occurred in BSI episodes caused by Gram negative episodes (70.3%%) versus 33.3% in Gram positive. Table 4 describes where the interventions occurred.
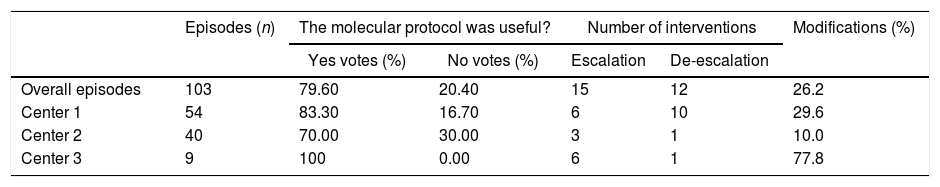
Clinical evaluation of the molecular protocol and therapeutic interventions performed on BSIs episodes.
| Episodes (n) | The molecular protocol was useful? | Number of interventions | Modifications (%) | |||
|---|---|---|---|---|---|---|
| Yes votes (%) | No votes (%) | Escalation | De-escalation | |||
| Overall episodes | 103 | 79.60 | 20.40 | 15 | 12 | 26.2 |
| Center 1 | 54 | 83.30 | 16.70 | 6 | 10 | 29.6 |
| Center 2 | 40 | 70.00 | 30.00 | 3 | 1 | 10.0 |
| Center 3 | 9 | 100 | 0.00 | 6 | 1 | 77.8 |
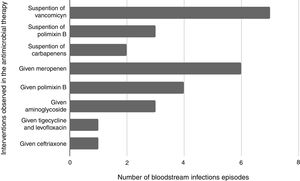
De-escalation to targeted therapy was possible in 12 (11.6%) of the episodes, including suspension of polymyxin B (n=3), carbapenem (n=2), vancomycin (n=7). Escalation occurred in fifteen episodes, most frequently in those with an indication of carbapenem (Fig. 1). Antimicrobial therapy was inadequate in 12% (3/25) of the modified interventions. Among BSI episodes in which the patients received no interventions after the optimized protocol results were reported, antimicrobial therapy was inadequate in 10.2% (8/78) of the episodes.
DiscussionThe emergence of rapid diagnostic technologies, including MALDI-TOF MS and genetic assays, have dramatically reduced the time to identify microorganisms from blood samples.20,27-29 Use of these tests is an increasingly important component of antimicrobial stewardship service and routine clinical microbiology.30 Numerous strategies can be used to identify pathogens from positive blood samples by MALDI–TOF MS31–35 and the accuracy of these methods has been reported, but further investigation into TAT and the clinical importance of this technology for pathogen identification had yet to be performed.
Herein we have combined MALDI-TOF MS and in-house real-time PCR for performing pathogen identification and susceptibility testing directly from BC bottles using samples from 103 episodes of BSI in 91 patients. The average time to report bacterial identification and antibiotic resistance results to the clinical team was 35h earlier with the developed protocol compared to conventional techniques. Identification at the genus level using the rapid protocol was 93.5% in agreement with the results from conventional identification methods.
Vlek et al. conducted a hybrid investigation of 218 patients with bacteremia in which MALDI-TOF was utilized for bacterial identification and the results were compared with those from conventional testing. The average time to bacterial identification was 28.8h earlier with MALDI-TOF compared to conventional testing (16.4 versus 45.2h); earlier pathogen identification enhanced the ability of the physicians to prescribe appropriate treatment.36
In this study, 27 interventions were performed with 22 successful antimicrobial therapy changes. Modifications occurred most frequently in Gram negative episodes (70.3%% versus 33.3% in Gram positive). Clerc et al. reported an observational investigation in which MALDI-TOF results (without stewardship mediation) were associated with antimicrobial treatment change in 35.1% of patients with Gram negative bacteremia.22
Successful interventions in the present study included three episodes in which the patients were receiving meropenem based on the morning Gram stain result, but their treatment was changed to polymixin B after results of the rapid protocol testing were received.
One patient had a previous BSI with KPC-producing K. pneumoniae and was receiving a combined therapy consisting of meropenem and amikacin. Molecular results were available 5.15h after Gram staining and reported the presence of both the KPC and rmtG methylase genes. Antibiotic therapy was modified to polymixin B, tigecycline, and ceftazidme/avibactam 41h before final susceptibility testing. The patient was neutropenic with acute lymphoid leukemia who, after adequate intervention and pathogen clearance, was able to undergo a bone marrow transplant.
Although good agreement for antimicrobial susceptibility testing between phenotypic (conventional) and optimized methods was found in this study, some of the interventions started based on molecular antimicrobial susceptibility results led to inadequate therapy. In one BSI episode, false-negative ESBL testing resulted in de-escalation of meropenem for cefepime which had to be re-escalated to carbapenem after the final susceptibility results were received. Two false-positive ESBL samples (tested positive for blaTEM by real-time PCR) resulted in the prescription of meropenem for cephalosporin susceptible pathogens.
A unique aspect of this study was that we used real-time reporting of the optimized protocol results directly to infectious diseases physicians so they could initiate immediate antimicrobial therapy evaluation and subsequent interventions when necessary. Previous studies have suggested that implementation of rapid diagnosis without real-time reporting to, and real-time feedback from, an antimicrobial stewardship team has little or no impact on patient outcomes.32,37,38 A few studies30,39 that reported findings from BSIs caused by Gram negative bacteria tentatively showed that fast pathogen identification and antimicrobial stewardship decreases in-hospital length of stay and aggregate expenditures.
In this study, we propose the use of a protocol which utilizes molecular and phenotypic techniques to rapidly identify bacteria and determine antimicrobial resistance directly from blood culture bottles in a diverse patient population. We did not evaluate the effect of this protocol on length of hospital stay, costs, impact on mortality, and other clinical outcomes compared with conventional methods for bacterial identification and antibiotic resistance testing. Further investigation should be conducted to better understand the performance and clinical impact of our proposed optimized protocol.
ConclusionsIn this study the use of a combined protocol of direct MALDI-TOF MS and real-time PCR yielded rapid and reliable bacterial identification and antimicrobial profiling in bacteremia episodes. This method gives reliable results to physicians 35h (median) earlier than conventional testing. In addition, we observed that several factors influence the clinical outcome of rapid diagnosis of BSI, including the interpretation of optimized results by the attending physician, target population, and clinical condition of patients. Use of an optimized protocol, such as the one reported in this study, requires rapid and adequate clinical support to effectively bring benefit to clinical treatment and rationalization of antimicrobial use.
Ethics approval and consent to participateThis study was approved by the Clinical Research Ethics Committee of the Federal University of São Paulo (Universidade Federal de São Paulo). All subjects read and signed informed consent forms for participation in the study.
FundingThis study was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior – Brasil (CAPES).
Conflicts of interestThe authors declare no conflicts of interest.
We thank Sarah Bubeck, Ph.D., from Edanz Group (www.edanzediting.com/ac), for editing a draft of this manuscript.