Vancomycin-resistant Enterococcus faecium (VREfm) has emerged as an important global nosocomial pathogen, and this trend is associated with the spread of high-risk clones. Here, we determined the genetic and phenotypic features of 93 VREfm isolates that were obtained from patients in 13 hospitals in Vitória, Espírito Santo, Brazil, during 2012–2013. All the isolates were vancomycin-resistant and harbored the vanA gene. Only 6 (6.5%) of the VREfm isolates showed the ability to form biofilm. The 93 isolates analyzed belong to a single pulsed-field gel electrophoresis lineage and presented six subtypes. MLST genotyping showed that all VREfm belonged to ST412 (the high-risk clone, hospital-adapted). The present study describes the dissemination of ST412 clone in the local hospitals. The clonal spread of these ST412 isolates in the area we analyzed as well as other hospitals in southeastern Brazil supports the importance of identifying and controlling the presence of these microorganisms in health care-related services.
Within the last two decades, Enterococcus faecium has emerged as an important global healthcare-associated pathogen because of its ability to colonize and cause disease in high-risk patients.1,2 This emergence can be explained in part as a result of the resistance of E. faecium to several antimicrobial agents, both intrinsic and acquired.3 Vancomycin is usually required for treatment, especially for invasive infections. However, data from nosocomial infection surveillance worldwide has revealed a growing percentage of vancomycin-resistant E. faecium (VREfm) clones.4,5
Antimicrobial resistance, together with virulence factors, contribute to the development of human enterococcal infections.6 Biofilm production has an important role in the pathogenesis of bacterial infections, since this feature allows the permanency of the microorganism by protecting it from the host defense mechanisms and can facilitate horizontal gene transfer, thereby contributing to antimicrobial resistance spreading.7,8
Changes in the epidemiology of E. faecium infections have been associated with the global dissemination of the high-risk clones that have acquired adaptive elements such as antimicrobial resistance and virulence genes, particularly in a defined subpopulation of E. faecium that is enriched in hospital isolates.1,2
The aim of the present study was to characterize 93 VREfm isolates obtained between 2012 and 2013 from various clinical specimens: catheter (1), wound (3), blood (4), urine (19), and rectal swab (66) of different patients at 13 hospitals in Vitória, Espírito Santo, located in southeastern Brazil, to increase the knowledge of the molecular epidemiological characteristics of this nosocomial pathogen.
The isolates were previously identified as VREfm using the Vitek2® system (bioMérieux, France) and confirmed by PCR, as described by Depardieu et al.9 The minimum inhibitory concentration (MIC) for vancomycin was determined using Etest (bioMérieux).
Biofilm formation was measured in 96-well polystyrene microtiter plates (Costar, USA), following 24h of incubation at 35°C, and staining with crystal violet as previously described by Stepanovic et al.10 The optical density (OD) of each crystal violet-stained well was measured at 570nm (TP-Reader Spectrophotometer, Thermo Plate, China). Enterococcus faecalis ATCC 29212 and Staphylococcus aureus 111711 were used as negative and positive controls, respectively. All tests were carried out in experimental and biological triplicates. Based on the bacterial biofilm OD, isolates were classified into four categories: non-biofilm producer, weak, moderate, or strong biofilm producer. The cutoff OD (ODc) was defined as three standard derivations above the mean OD of the negative control. Isolates were classified as follows: OD<ODc=non-biofilm producer; ODc<OD<2ODc=weak biofilm producer; 2ODc<OD<(4ODc)=moderate biofilm producer; and OD>4ODc=strong biofilm producer.
Genomic DNA from E. faecium was extracted following the method described by thermal lysis and used as template for multiplex PCR of the van genes, as previously described by Depardieu et al.9
Pulsed-field gel electrophoresis (PFGE) was performed after macrorestriction with SmaI in a CHEF-DRIII system (Bio-Rad, USA), according to Saeedi et al.,12 and analyzed with Bionumerics v6.5 (Applied Maths, Belgium) using the unweighted pair-group method with the arithmetic mean (UPGMA) applying the Dice correlation coefficient. Isolates were designated as the same pulsotype if they shared at least an 80% similarity in the band patterns and the same subtype if showed an identical band pattern.
One isolate of each subtype was characterized by MLST method according to the recommendations described in the E. faecium MLST database (http://efaecium.mlst.net/).
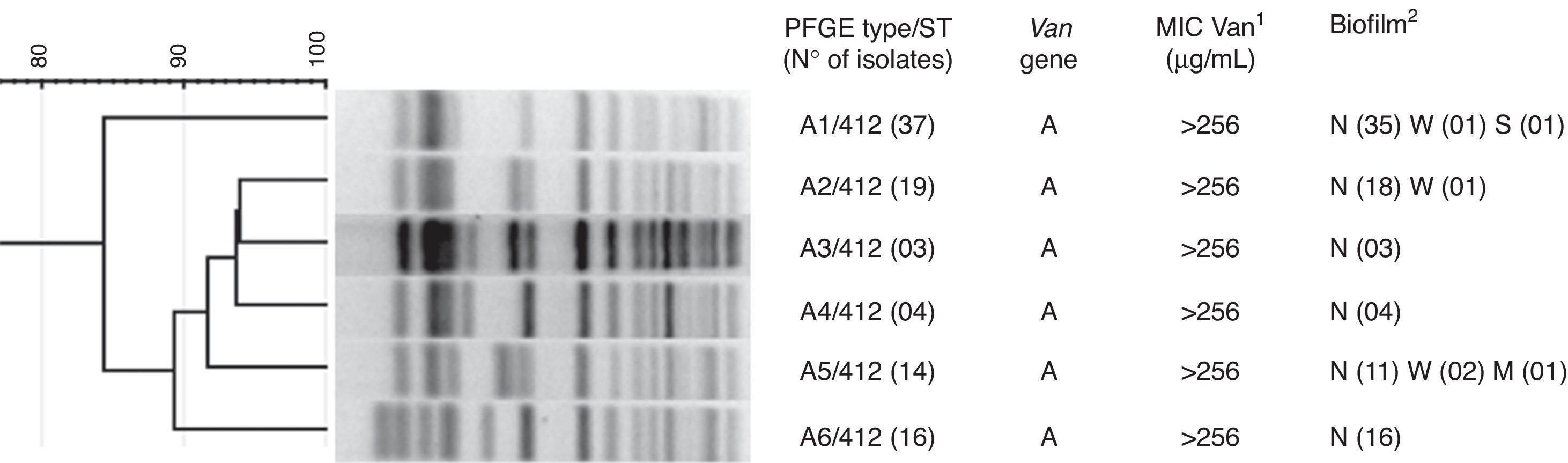
All isolates harbored the vanA gene and showed vancomycin MICs>256μg/mL (Fig. 1). In addition, 87 (93.5%) of the isolates could not form biofilm on a polystyrene surface. However, four (4.3%) of the isolates showed a weak ability to produce a biofilm, 1(1.1%) presented a moderate biofilm-forming ability, and the remaining strain presented strong biofilm production.
The 93 isolates belonged to the same PFGE lineage and presented six subtypes (A1-A6) (Fig. 1). The MLST results showed that all VREfm subtypes of lineage A presented ST412.
Our study describes the microbiological and epidemiological characteristics of VREfm clinical isolates that were obtained in different hospitals in a southeastern region of Brazil. The vanA- and vanB-mediated glycopeptide resistance occurs frequently in VREfm, and both types are carried out by transposons (Tn1546 and Tn1547, respectively).13 In the present study, vancomycin resistance was common in all isolates, and they all harbored the vanA gene. This result is consistent with the vancomycin-resistance phenotype, as all isolates were high-level resistant to vancomycin. In Brazil, clinical studies after outbreaks in different states have reported the emergence and prevalence of VREfm isolates carrying the vanA gene.6,14 The widespread prevalence of the vanA gene in E. faecium has also been observed in Canadian and European studies.2,5 The formation of multilayered biofilm in enterococci is a complex and multifactorial process.15 Studies on E. faecium isolates have shown the low or moderate ability of this species to form biofilm.15,16 In the present study, biofilm formation was observed in only six (6.5%) isolates. Paganelli et al.16 observed that E. faecium strains of different phylogenetic clades form biofilm with distinct properties and suggested that under different ecological conditions, different types of biofilms are produced, possibly contributing to adaptation to different niches.
PFGE profiles and MLST data indicate that there is a clonal dispersion among the VREfm clinical isolates analyzed in the present study. The strains showed a homogeneous pattern that was associated with a conserved presence of the vanA gene for all isolates. Notably, the 93 VREfm were clustered into only one lineage, the ST412, which belongs to the high-risk clones complex. Molecular epidemiological studies have shown the global spread of high-risk clones, which is associated with the majority of nosocomial outbreaks and clinical infections in all continents. The wide distribution of specific subpopulations seems to have been facilitated by the cumulative acquisition of antibiotic resistance, virulence characteristics, and the ability to acquire different genetic elements via horizontal transfer.14
Damani et al.17 described ST412, which belongs to clonal complex 17 (CC17), for the first time during an epidemiological study of VREfm isolates from Greece in 2010, where this clone was predominant in Greek clinical settings.
VREfm isolates belonging to CC17 are predominant in sporadic cases and during outbreaks in Brazil. Studies have shown that ST412 was the most frequent sequence type in hospital environments in Brazil and four other countries in South America, including Colombia, Ecuador, Peru, and Venezuela.6,18,19 The emergence of VREfm ST412 has been observed in southeastern Brazil, indicating a strong correlation between this strain and the hospital environment.6,19,20 The present study found that ST412 is a well-established clone in hospitals of Vitória, Espírito Santo, Brazil. The clonal spread of ST412 among hospitals in different areas of the country indicates an inter-hospital spread and emphasizes the need for the application of stringent control measures to decrease the risk of dissemination of the bacteria, such as the isolation of infected patients, increased environmental cleaning, and improved antimicrobial therapy.
Ethics statementThe present research received ethical and methodological approval from the Research Ethics Committee of the Center of Health Sciences of the Universidade Federal do Espírito Santo (Protocol 65/2011).
Financial supportThis work was supported by Fundação de Amparo à Pesquisa do Estado do Espírito Santo (FAPES) and Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq).
Conflicts of interestThe authors declare no conflicts of interest.
We acknowledge the contribution of PhD Nazareth Magnago Klein (Federal University of Espírito Santo) for the supply of the bacterial strains and PhD Thiago César Nascimento (Federal University of Juiz de Fora) for the data analysis.