Hepatitis C virus (HCV) is the leading cause of liver diseases including cirrhosis and hepatocellular carcinoma and it affects an estimated 3% of the world's population which corresponds to over 130 million HCV positive patients worldwide.1 HCV is an enveloped virus with a positive single stranded RNA genome in the family Flaviviridae. Due to its RNA genome, HCV shows substantial nucleotide sequence variability with nucleotide substitution rate of 1.44×10−3 per site/year resulting in evolution into seven major genotypes including multiple subtypes and quasispecies.2 RNA viruses evolve rapidly2 and changes in epidemiological patterns have been reported from different regions of the world. Several studies have reported HCV genotypes distribution in Khyber Pakhtunkhwa (KP), Pakistan,1,3,4 while none has so far reported the changing epidemiological pattern of HCV genotypes in the province. The aim of this study was to find out the frequency and existing pattern of HCV genotypes distribution in KP by a modified genotyping assay and sequencing.
A total of 510 HCV infected patients who were registered at various healthcare units at Peshawar city for free treatment or diagnosis under the prime minister's Hepatitis C control program were included in this study as per their consent and ethical approval of the Institutional committee. Qualitative detection of HCV and HCV genotyping was carried out by a modified reverse transcription-polymerase chain reaction (RT-PCR). Sequencing of the core gene was used for genotype authentication.
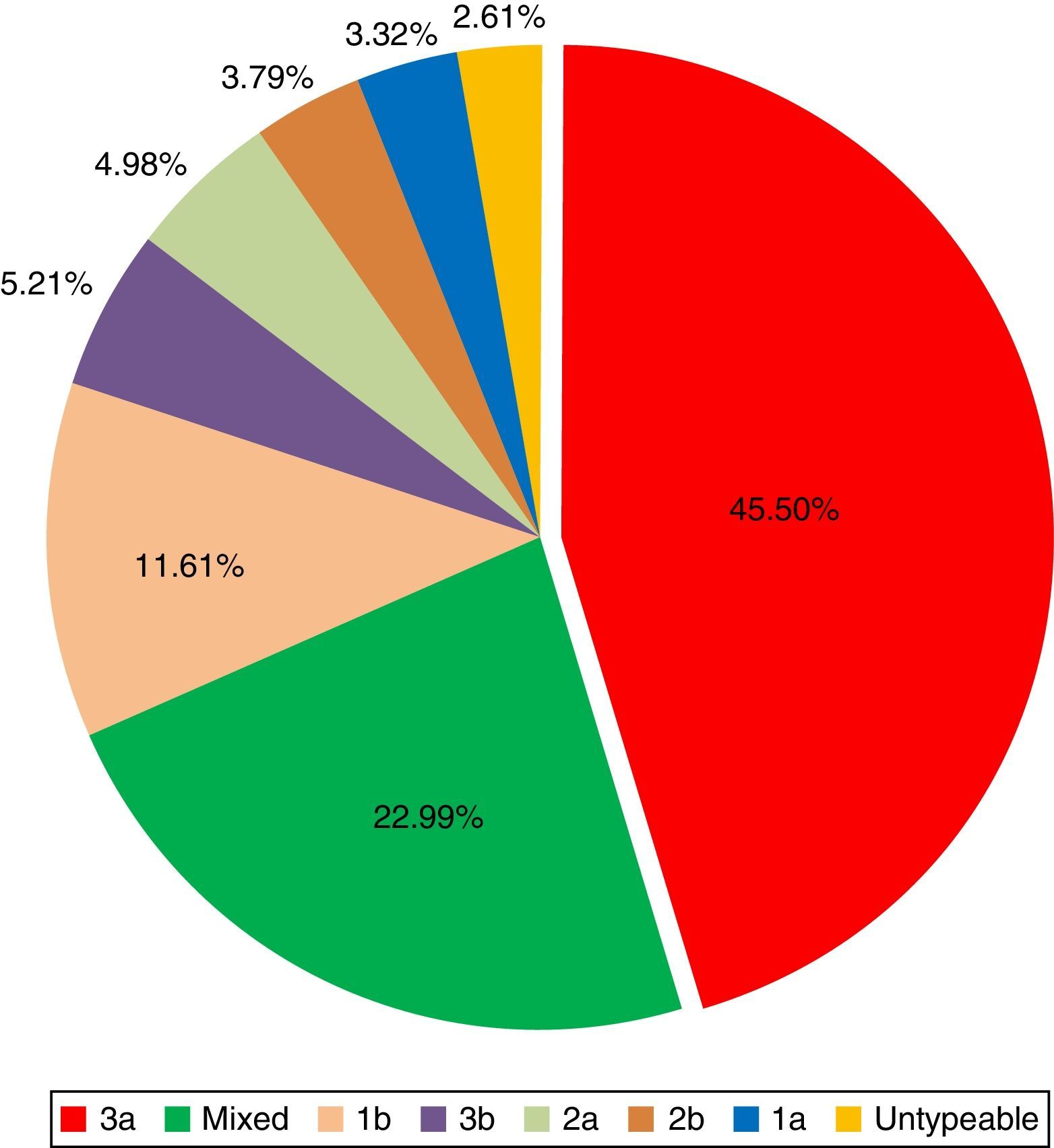
Out of the total, active HCV infection was detected in 422 (82.75%) patients. HCV 3a (45.5%) was found to be the most abundant subtype followed by a high proportion of mixed genotypes infection (22.99%). HCV 1b accounted for infection among 11.61% of the patients while 3b was detected in 5.21% of the cases. Rare genotypes or subtypes encountered were 2a, 2b, and 1a. Among mixed genotypes, 3a/1b was the most prevalent (49%) followed by an equal distribution of 3a/3b and 1a/1b (15%). Similarly, 1b/3b, 1b/2a, and 1a/3b mixed genotypes were detected in 9%, 6%, and 4% of the patients, respectively.
Previous studies have reported a much higher prevalence of HCV 3a (ranging from 70 to 90%) both in KP3,4 and Pakistan,5 while in this study only 45.5% of individuals were infected by HCV 3a. One of the most important findings in this study was the presence of significantly high proportion of subtype 1b and mixed infections than the previously reported frequencies for these genotypes. The existing prevalence of other subtypes such as HCV 3b (5.21%), 2a (4.98%), 2b (3.79%), and 1a (3.32%) also revealed a very different pattern from those reported earlier.3–5 A study reported subtype 1a and 1b to be present in an equal proportion of 7.7% in Peshawar district of KP.3 In contrast, we found higher prevalence of subtype 1b compared to subtype 1a (Fig. 1). Possible explanation for the observed shift in genotype pattern distribution in KP could be the high rate of sequence evolution or natural HCV recombination, inadequate sensitivity of old genotyping assays to detect the correct type, or the migration of people from Tribal regions and bordering Afghanistan due to the ongoing war.
Among mixed genotypes, 3a/1b was found to be the most prevalent pattern (49%) followed by an equal distribution (15% each) of 3a/3b and 1a/1b, and other mixes (Fig. 1). Being RNA viruses, HCV types do not exhibit cross immunity and mixed infections have been reported from KP and Pakistan.1,3–5 HCV subtype 1b and 3a were the two commonest subtypes present either together or in combination with other subtypes which reinforces our previous observation about the most prevalent HCV types in KP. The proportion of patients infected with subtypes 2a and 2b has gradually decreased. This can partly be attributed to the good response rates seen for these particular genotypes in KP and the effectiveness of the control strategies against these subtypes.
This study concludes a change in the pattern of HCV genotypes distribution in KP province of Pakistan with HCV subtype 1b and mixed genotype infections replacing IFN-responsive HCV strains. The newly emerging pattern of HCV genotypes could have consequences for hepatitis C control and management strategies.
Conflicts of interestThe authors declare no conflicts of interest.
Authors acknowledge support of the Directorate of Science and Technology KP, for the study [Project No. Dirtt: S&T/KP/PRS/FINANCIAL ASSISTANCE/2011-12].