The methicillin-resistant Staphylococcus aureus (MRSA) USA300-Latin American variant (USA300-LV) lineage is well documented in northern Latin American countries. It has replaced established clones in hospital environments. We herein report a systemic infection caused by a USA300-LV isolate in a 15-year-old boy, from a low-income area of Rio de Janeiro, previously colonized by the same strain. During hospital stay, seven pvl-positive MRSA USA300-LV isolates were recovered by nasal swab, blood and abscess secretion. The patient underwent intravenous vancomycin, daptomycin, and oral sulfamethoxazole/trimethoprim, and was discharged after 45 days after full recovery. This is the first documented case of a community-acquired MRSA infection caused by the USA300-LV variant in Brazil in a previously colonized adolescent with no history of recent travel outside of Rio de Janeiro. The need for improved surveillance programs to detect MRSA colonization in order to control the spread of hypervirulent lineages among community and hospital settings is highlighted.
Staphylococcus aureus colonizes about 30% of human nares. Such colonization is an important source of person-to-person bacterial transmission, especially in low socioeconomic status communities, characterized by a large number of subnormal agglomerates.1,2 Recent studies have proposed that previous nasal colonization with methicillin-resistant S. aureus (MRSA) strains could increase the risk of a subsequent staphylococcal infection, such as skin and soft tissue infections (SSTI), pneumonia, osteomyelitis, bloodstream infections (BSI) and endocarditis.1,3
Community-associated MRSA (CA-MRSA) USA300/ST8/SCCmecIVa strains are considered epidemic and widely disseminated in North America as a causative agent of SSTI.4 In general, CA-MRSA strains are known for their ability to carry a wide range of virulence determinants, such as the Panton-Valentine leucocidin (PVL) gene (pvl), which has been associated with invasive and severe diseases, particularly in children.5 However, differently from North America, a USA300-related genetic lineage (designated USA300 Latin American variant [USA300-LV]) emerged and has become epidemic in countries of the northern region of South America, such as Colombia, Venezuela, and Ecuador, as the causative agents of bloodstream infections. It has replaced the hospital-associated MRSA clone designated “Chilean/Cordobes”.6–9 The first description of USA300-LV was in 2009 in a prospective multicenter study that aimed to evaluate the molecular epidemiology of MRSA isolates recovered from 32 high-level care hospitals distributed in four Latin American countries in 2006 and 2008.9 Similarly to other CA-MRSA strains, USA300-LV isolates can harbor the PVL gene; however, unlikely USA300/ST8/SCCmecIVa, the USA300-LV lacks the arginine catabolic mobile island element (ACME), and most USA300-LV isolates carry different variants of type IV methicillin resistance cassette.6,9
In Rio de Janeiro, Brazil, hospital-associated MRSA (HA-MRSA) USA100/ST5/SCCmecII and CA-MRSA USA1100/ST30/SCCmecIV lineages are frequently associated with BSI and SSTI, respectively.10,11 However, in this study, we aimed to describe for the first time in Brazil the occurrence of a systemic infection caused by a CA-MRSA isolate belonging to the pvl-positive USA300-LV/SCCmecIVd/t008 lineage in the nares of a Brazilian 15-year-old boy colonized by the same MRSA strain.
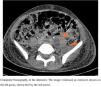
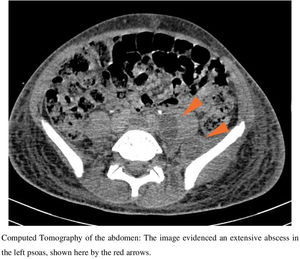
Case reportOn October 19th, 2019, a 15-year-old boy from a low-income neighborhood of Rio de Janeiro, with no previous history of travel outside Brazil and with no comorbidity, entered a Brazilian university hospital with a two-week history of lumbar pain and polyarthralgia after a body trauma during a soccer game. Upon admission, the patient was febrile and presented edema, redness, and heat in right elbow and left shoulder, suggesting a disseminated and complicated S. aureus SSTI. Computed tomography (CT) of the abdomen evidenced an extensive abscess in left psoas (Fig. 1) and peripheral pulmonary embolic lesions, with no documentation of endocarditis in the transesophageal echocardiogram. Also, a magnetic resonance imaging (MRI) of the spine showed a posterior compression from L2 to S2, which was suggestive of a subdural empyema in the sacral region extending to the lumbar spine. A screening nasal swab for MRSA colonization was performed, and blood was drawn. Antimicrobial therapy began immediately. Only in the eleventh day of hospitalization the infection was controlled, with clinical improvement and serial negative blood cultures. The patient received intravenous vancomycin (15-20 mg/kg/day depending on serum levels of vancomycin), followed by daptomycin (10-12 mg/kg/day), switching to oral sulfamethoxazole/trimethoprim (400/80 mg/q 6 h) up to six months.
All blood cultures were processed using the automated BactAlert system (BioMerieux, Durham, NC, USA). After determining positivity, blood was cultured on sheep blood agar (Plast Labor, Rio de Janeiro, Brazil) for 24 hours with incubation at 37° C, in addition to abscess drainage collection. Bacterial colonies were identified using the VITEK®2 (BioMerieux, Durham, NC, USA) automated identification and susceptibility testing system. All isolates recovered (blood, psoas, and epidural abscess) revealed growth of MRSA. Also, the nasal swab was seeded on CHROMagar MRSA (Plast Labor, Rio de Janeiro, Brazil), and a MRSA isolate was confirmed by VITEK®2 (BioMerieux, Durham, NC, USA).
Seven MRSA isolates were sent to a reference laboratory. All were confirmed to be S. aureus by MALDI-TOF MS (matrix-assisted laser desorption/ionization-time of flight mass spectrometry) and methicillin-resistant by cefoxitin disk (CLSI, 2019). The isolates were subjected to disk-diffusion and broth microdilution test to determine the Minimal Inhibitory Concentration (MIC) for common antimicrobials (CLSI, 2019). In order to describe the SCCmec type, the presence of pvl, the pulsed field gel electrophoresis (PFGE) pattern, and the Sequence Type (ST), all isolates were processed as described by Chamon et al.11 In addition, the isolates were also characterized as for their SCCmec subtype and spa typing, and the presence of ACME was determined by polymerase chain reaction (PCR).12–14
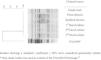
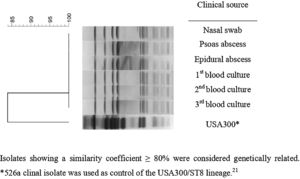
All MRSA isolates were susceptible to chloramphenicol, ciprofloxacin, clindamycin, erythromycin, gentamicin, rifampicin, sulfamethoxazole-trimethoprim and tetracycline. All presented the same MIC values of 64 μg/mL, 0.5 μg/mL, 1 μg/mL, and 1 μg/mL for oxacillin, daptomycin, linezolid, and vancomycin, respectively. All isolates presented the SCCmec type IVd and the pvl gene but were negative for ACME operon. PFGE revealed that all isolates presented the same band patterns (Fig. 2) and all were related to ST8/CC8, suggesting that all belonged to the USA300 lineage. However, the band pattern observed was different from those observed for a USA300 control strain (526a),11,21 and the spa type was thus determined as t008. Therefore, considering the data, we could characterize all isolates as belonging to the CA-MRSA pvl-positive ACME-negative USA300-LV/ST8/SCCmecIVd lineage.
DiscussionStaphylococcus aureus is an important pathogen in hospital and community environments. It is also a member of the skin microbiota of approximately 30% of the population. Its colonization is considered a risk factor for the development of invasive infections among hospitalized patients.1,3 Kim et al. conducted a meta-analysis and reported that subjects from community settings previously colonized with S. aureus, specially by MRSA isolates, were also associated with a greater risk of having a S. aureus infection.3 Also, Neves et al. suggested that children from a middle/low socioeconomic status were more likely to be colonized by MRSA strains.2
Although the incidence of CA-MRSA infections in Brazil is considered low (< 10%), recent studies have described an increase in the occurrence of CA-MRSA SSTI among pediatric patients.15,16 In the present report, a 15-year-old boy from a low-income area of Rio de Janeiro without any known risk factor for MRSA infection was admitted at our university hospital presenting clinical signs and symptoms suggestive of a disseminate staphylococcal disease. It is noteworthy that the boy, besides being infected, was also colonized by a MRSA isolate.
The microbiologic analysis showed that all seven MRSA isolates recovered presented the same PFGE pattern. All belonged to the CA-MRSA pvl-positive ACME-negative USA300-LV/ST8/SCCmecIVd/t008 lineage, described here for the first time in Brazil. The most important genetic distinction between the USA300-LV and North American USA300 lineages is the absence of the ACME island in USA300-LV, which is replaced for a gene cluster-encoding proteins involved in the metabolism of copper and mercury.6,17,18 However, similarly to the USA300 epidemic clone from North America, USA300-LV variants can cause invasive infections in healthy individuals, as shown in the present case report.
The USA300-LV lineage is a parallel USA300 epidemic clone that affects predominantly countries of the northern region of South America. It is frequently associated with community-acquired SSTI but also with nosocomial BSI.6–9,17 Recently, Arias et al. described that USA300-LV isolates can completely replace prevalent hospital-associated clones in Colombia and Ecuador.8 Moreover, this lineage was already described in European countries, such as Germany and Switzerland, however isolated from patients traveling in or from South America.18,19 Interestingly, Takadama et al. suggested that a PVL-positive CA-MRSA USA300-LV clone has evolved and disseminated in Japan, giving rise to a new clone called USA300-LV/J. This shows the adaptability capacity of such lineage to disseminate to distinct geographic regions.20 In the present study, we have described a systemic infection caused by the USA300-LV lineage in a healthy boy with no previous history of travelling outside Brazil. He was previously colonized by this CA-MRSA clone, suggesting a community acquisition of this strain. It is important to mention that the community transmission of the USA300-LV variant has already been reported in Germany.19
ConclusionAs Brazil is a touristic route for many global travelers, we emphasize the possibility of a community spread of a hypervirulent CA-MRSA lineage, such as the USA300-LV, which is widely adapted to and disseminated in some Latin American countries, among them Brazil. Therefore, the improvement of surveillance programs and financial support for epidemiological studies on the community colonization by MRSA isolates, together with antimicrobial resistance studies, are urgent and necessary in order to prevent the dissemination of hypervirulent MRSA isolates into hospital and community environments in Brazil.
Ethics approval and consent to participateThe present case report was approved by the “Comitê de Ética em Pesquisa da Faculdade de Medicina da Universidade Federal Fluminense” under CAAE no. 51559821.3.0000.5243. A written informed consent was obtained from the patient's mother, who authorized the publication of this case report and all accompanying images.